 There would be 20 40 options, or a mind-boggling 1. Thus, the total number of resulting tripeptides possible would be 20 X 20 X 20 or 20 3, or 8,000 different tripeptide sequences! Now think about how many options there would be for a small peptide containing 40 amino acids. At each position, there are 20 different options that can be incorporated. So the cutoff of 50 amino acids (MW about 5500) derived from open reading frames seems like the best arbitrary cutoff going from peptides to proteins.ĭue to the large pool of amino acids that can be incorporated at each position within the protein, there are billions of different possible protein combinations that can be used to create novel protein structures! For example, think about a tripeptide made from this amino acid pool. As we learned in the Chapter 2, hydrogen bonds are directional (exhibiting a bond angle of 180) therefore, the peptide backbone must adopt specific conformations in order for atoms in peptide bonds to hydrogen bond with each other. Even this ignores small proteins that have been isolated and characterized from cells. Submissions of DNA sequences to the NIH GenBank must encode proteins no smaller than about 66 amino acids (MW about 7250). The most simple and elegant arrangement is a right-handed spiral conformation known as the alpha-helix. If no cutoff was used, the number of proteins encoded by the genome would be huge. Pauling and Corey twisted models of polypeptides to find ways of getting the backbone into regular conformations which would agree with alpha-keratin fibre diffraction data. For the yeast genome, annotated proteins were defined as those made from an open reading frame leading to a polypeptide of 100 amino acids (which on average has a molecular weight of 11,000. derived from proteins by proteolysis, and/orĪs genomes were sequences and annotated, some arbitrary parameters were set."intrinsically disordered" with no definite fold,.Peptides, on the other hand, could be structures that are: Storz et al consider polypeptides to be small proteins if they are encoded in the genome by a continuous stretch of DNA base in an "open reading frame" (doi: 10.1146/annurev-biochem-070611-102400). One way to differentiate them is by how they are synthesized in vivo. Others suggest that structures with 40-50 amino acids should be considered small proteins. The human version has over 34,000 amino acids and a molecular weight > 3.9 million! Some consider structures with fewer than 50 amino acids to be p eptides (Figure 2.13). 
Proteins range in size from around 50 amino acids in length to the largest known protein, titin (aka called connectin), a muscle protein. 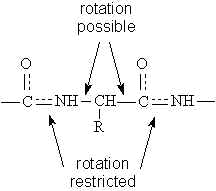 
The order and nature of amino acids in the primary sequence of a protein determine the folding pattern of the protein based on the surrounding environment of the protein \): Peptides and Proteins are macromolecules built from long chains of amino acids joined together through amide linkages. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
